Thoughtworks Engineering for Research (E4R) is our initiative to advance scientific research by working with leading global scientific organizations on their challenging computational problems.
This blog is an overview of one such project that began in mid-2019 and in early 2020 during the pandemic, was further guided by Dr. Gautam Menon, Professor of Physics and Biology at Ashoka University, India.
EpiRust. The beginning.
The COVID-19 pandemic established that infectious diseases are still a global threat. An epidemic affects not only people’s health but also the economy and society's morale at large. COVID-19's accelerated spread motivated the public health policy makers to better understand the virus and its characteristics that would help discover interventions to arrest its spread.
To analyze and predict a pandemic such as COVID-19's behavior is a complex task. Pandemics are rare and have few historical references. The last pandemic that is similar to what is happenig now, occurred over a century ago and that era's data is so thin and barely useful. Additionally, every pandemic could have numerous unique and local characteristics – based on demographics, local climate and the disease itself.
As a pandemic starts unfolding, public health policy makers have limited time to assess the risk and prepare a response strategy. Such a response strategy must adapt to an evolving situation, involve various what-if and if-what scenarios.
Historically, policy makers relied on their own experience, skills and imagination to describe these scenarios. Such descriptions were often implicit and informal. Today, computer simulations help bridge this gap and aid in the development of response strategies. They perform what-if experiments to understand the spread of new pathogens and optimize interventions based on available resources. The success of such simulations lies in the right choice of modeling approaches and their scale.
Our simulation, that we named – EpiRust allows a researcher to introduce a pathogen into a virtual society and observe the disease’s spread under various circumstances – when people go to work, under lockdowns and so on.
Why an agent-based model?
Simulation as a technique has existed for decades – to evaluate what-if and if-what scenarios in policy making. For epidemic modeling, researchers have used differential equations for multiple use cases from simple to complex.
These models are computationally less intensive and run faster. However, an epidemic emerges from interactions among individuals. And, as these models do not capture interactions among individuals, they cannot explain the spread of an epidemic. In addition, equation-based models also face challenges with interventions.
An alternative to differential equations based models are agent-based models. Here, agents represent people in a simulated society, living their daily lives. Thus, it is a bottom-up approach for modeling individual behaviors, their decisions and their interactions with others. This collectively helps in studying emerging dynamics such as spread of an epidemic.
One can simulate a virtual environment mimicking geography, climate, population, buildings and other representative information. For example, an individual could infect companions while traveling on a bus or studying at a school. A policy maker could subsequently propose interventions such as decongestion of public transport and closure of schools. Such detailed simulations are called 'microsimulations.'
Our motivation has been to develop a robust, performant, flexible framework for large-scale, agent-based epidemic simulation models. To harness the compute power of underlying hardware, we decided to use the Rust language for its performance and safety features. We named it EpiRust to reflect its epidemiological basis and the use of the Rust language for implementation.
Designing EpiRust
EpiRust is amongst the first large-scale, agent-based epidemic simulation frameworks developed in India. It follows a minimalist approach for modeling an epidemic in a virtual society along three main aspects – population, geography and disease.
An agent (a person in the population) is initialized into the virtual society based on whether they work or stay at home, whether they use public transport and more.
The model mimics a city’s geography by allocating places on a grid for home, workplace and transport. This shows how the rates of infection change with mobility.
All the agents are treated as susceptible to the disease. Their status changes when they are exposed to the infection, fall ill and either recover or succumb to it. The model also accounts for external interventions like lockdowns, isolation and vaccination.
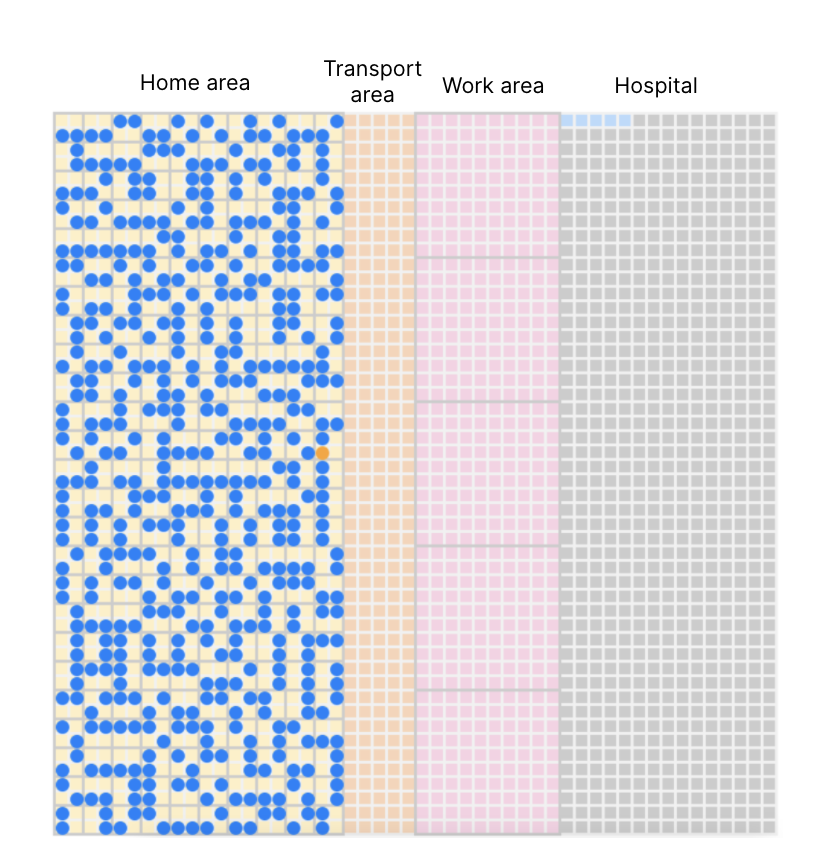

Grid-based geography similar to a minimalist planned city
Choosing tech that works
To implement large-scale agent-based simulations, developers historically relied on C and C++ for better performance. However, developers using these languages also face low productivity in terms of development time. Additionally, software quality issues emerge because of the complex explicit memory management.
While C#, Go, Java, Julia, and Python are designed for automated memory management via garbage collection. At the same time, garbage collection introduces its own overheads potentially resulting in overall lower performance.
Considering all of this, we chose the Rust language for its performance and built-in checks to ensure safer memory management even for concurrency, without a need for garbage collection.
Achieving robustness, flexibility and performance
EpiRust realized robustness by eliminating problems of memory management – Rust retained only one copy of the agents and related data. It also did not need complicated clean up processes.
Rust constructs, like modules, structures and traits, separated disease dynamics from other factors. This allowed different scenarios to be simulated without changing the source code, therefore making it flexible.
At this stage, we faced two hurdles – first, the overall outcome of the simulation was dependent on the order of agent evaluation for their next infection state and second, these simulations required large amounts of memory.
We resolved both issues by replacing data structures and algorithms with what was more suitable for grid-based geography. The former issue was dealt with by using double buffering – one for read-only state and another for write-only. For the latter issue, we chose spatial indexing.
What-if and if-what simulation
The next step was to deploy the city-model based on two scenarios:
The baseline, where the infection trajectory was traced during business as usual
The intervention, when mitigative measures were put in place
The second scenario brought another challenge – the model had to identify the costs of such measures vis-a-vis their benefits. It needed to answer questions: Is the reduction in infection justified, considering the negative effects of the lockdown?
The model also took into account eventualities like groups that would disobey restrictions, by adding them to the number of essential workers who would continue to work. We ensured accuracy by running the model at least five times for statistically stable results.
EpiRust implementation
We used EpiRust to simulate the population of the city of Pune in India – a city that spans 331 square kms and has more than 3 million residents – to test whether the model could replicate results at scale.
Parameter |
Comment |
Value |
number_of_agents |
Total number of agents to run simulation with |
3137224 |
grid_size |
Size of the grid (n*n) |
5660 |
working_percentage |
Percentage of working class population |
70% |
public_transport_percentage |
Percentage of working individuals who use public transport |
80% |
essential_workers_percentage |
Percentage of working individuals who are essential workers and will continue working even during lockdown |
10% |
lockdown_at_number_of_infections |
Number of infections that will impose lockdown |
100 |
hospital_beds_percentage |
Percentage of hospital beds per 1000 individuals |
0.003% |
starting_infections |
Number of infections at the beginning of the simulation |
32645 |
Input parameters
When implementing the COVID-19 model, we faced two major hurdles. The first – infering COVID-19 disease dynamics from the clinical data of patients. The second – evolving interventions in the model to match the pace of changes in government policies such as lockdown and isolation guidelines.
Our collaboration with Dr. Menon from the Ashoka University steered us to implement a more detailed disease dynamics model; the Mordecai Model which differentiates between asymptomatic/symptomatic, mild/severe infections, and hospitalization/non-hospitalization disease states.
Initial outcomes and future outlook
"Agent-based models provide granular and detailed approaches to disease spread, but combining complexity and scalability in computational implementations of such models is a permanent problem. The choice of the programming language Rust for the epidemiological simulator – EpiRust, is an inspired one as it provides a way of addressing this dual challenge. EpiRust provides its users both the flexibility and speed which other popular agent-based modeling codes, such as GAMA, struggle with." - Dr. Gautam Menon, Professor of Physics and Biology at Ashoka University, India.
Multiple runs of the simulation indicated that lockdowns were an effective means of controlling the disease’s spread. And, EpiRust proved to be a useful tool in the epidemiological study of COVID-19.
EpiRust is open sourced and available on GitHub. For more information, please read our paper, 'Framework for large-scale agent-based epidemiological simulations' from the SIMS2020 Conference. Our collective hope is to encourage more home-grown innovation in India.
The collaboration with Dr. Menon and his team at Ashoka University, on EpiRust, led to BharatSim – India’s first ultra large-scale agent-based simulation framework to model social dynamics and policy making related to epidemiology, economics and climate change mitigation. Its development is funded by the Bill and Melinda Gates Foundation.
Disclaimer: The statements and opinions expressed in this article are those of the author(s) and do not necessarily reflect the positions of Thoughtworks.


















